Efficient blockLASSO for Polygenic Scores with Applications to All of Us and UK Biobank
New paper!
https://www.medrxiv.org/content/10.1101/2024.06.25.24309482v1
Efficient blockLASSO for Polygenic Scores with Applications to All of Us and UK Biobank
Timothy G Raben, Louis Lello, Erik Widen, Stephen DH Hsu
Abstract: We develop a "block" LASSO (blockLASSO) method for training polygenic scores (PGS) and demonstrate its use in All of Us (AoU) and the UK Biobank (UKB). BlockLASSO utilizes the approximate block diagonal structure (due to chromosomal partition of the genome) of linkage disequilibrium (LD). LASSO optimization is performed chromosome by chromosome, which reduces computational complexity by orders of magnitude. The resulting predictors for each chromosome are combined using simple re-weighting techniques. We demonstrate that blockLASSO is generally as effective for training PGS as (global) LASSO and other approaches. This is shown for 11 different phenotypes, in two different biobanks, and across 5 different ancestry groups (African, American, East Asian, European, and South Asian). The block approach works for a wide variety of phenotypes. In the past, it has been shown that some phenotypes are more/less polygenic than others. Using sparse algorithms, an accurate PGS can be trained for type 1 diabetes (T1D) using 100 single nucleotide variants (SNVs). On the other extreme, a PGS for body mass index (BMI) would need more than 10k SNVs. blockLasso produces similar PGS for phenotypes while training with just a fraction of the variants per block. For example, within AoU (using only genetic information) block PGS for T1D (1,500 cases/113,297 controls) reaches an AUC of 0.63+-0.02 and for BMI (102,949 samples) a correlation of 0.21+-0.01. This is compared to a traditional global LASSO approach which finds for T1D an AUC 0.65+-0.03 and BMI a correlation 0.19+-0.03. Similar results are shown for a total of 11 phenotypes in both AoU and the UKB and applied to all 5 ancestry groups as defined via an Admixture analysis. In all cases the contribution from common covariates - age, sex assigned at birth, and principal components - are removed before training. This new block approach is more computationally efficient and scalable than global machine learning approaches. Genetic matrices are typically stored as memory mapped instances, but loading a million SNVs for a million participants can require 8TB of memory. Running a LASSO algorithm requires holding in memory at least two matrices this size. This requirement is so large that even large high performance computing clusters cannot perform these calculations. To circumvent this issue, most current analyses use subsets: e.g., taking a representative sample of participants and filtering SNVs via pruning and thresholding. High-end LASSO training uses ~ 500 GB of memory (e.g., ~ 400k samples and ~ 50k SNVs) and takes 12-24 hours to complete. In contrast, the block approach typically uses ~ 200x (2 orders of magnitude) less memory and runs in ~ 500x less time.
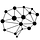
Our blockLASSO method for training polygenic scores (PGS) utilizes the approximate block diagonal structure (due to chromosomal partition of the genome) of SNP correlations. It reduces computational complexity by orders of magnitude: typically uses ~ 200x (2 orders of magnitude) less memory and runs in ~ 500x less time.
Summary of main PGS metrics for results in NIH All of Us (AoU), the UK Biobank trained with sets matching the size of those found in AoU (UKB∗ ), and for the UK Biobank using the maximum possible training sizes (UKB).
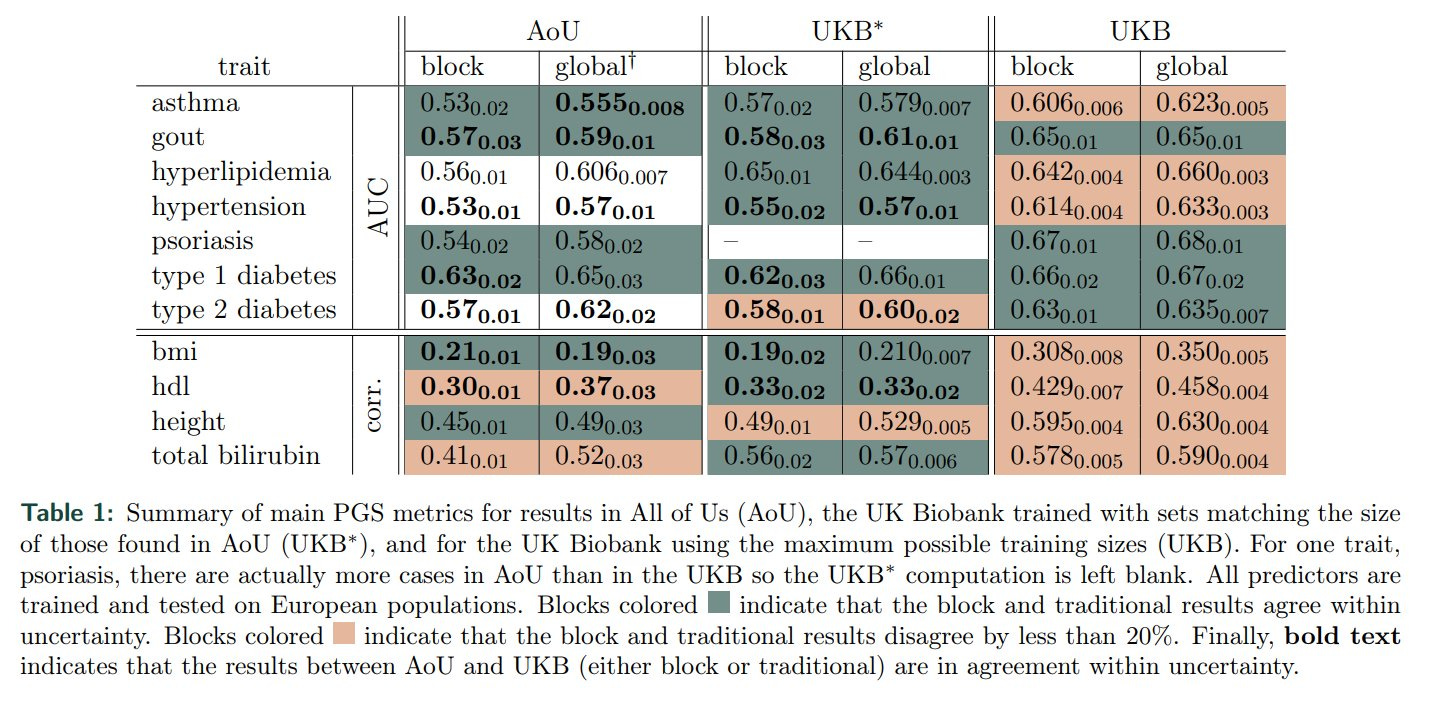
Note computational genomics is still DATA LIMITED! Algos are good enough - but significant gains will result from larger case populations (ie larger number of genotyped individuals with the condition).
Our earlier paper on this:
Biobank-scale methods and projections for sparse polygenic prediction from machine learning
https://www.nature.com/articles/s41598-023-37580-5
We especially need larger biobanks for non-European ancestry groups.
"For most case-control conditions, except T1D [in Euro populations], there are large gains that can be found from increased training sizes."
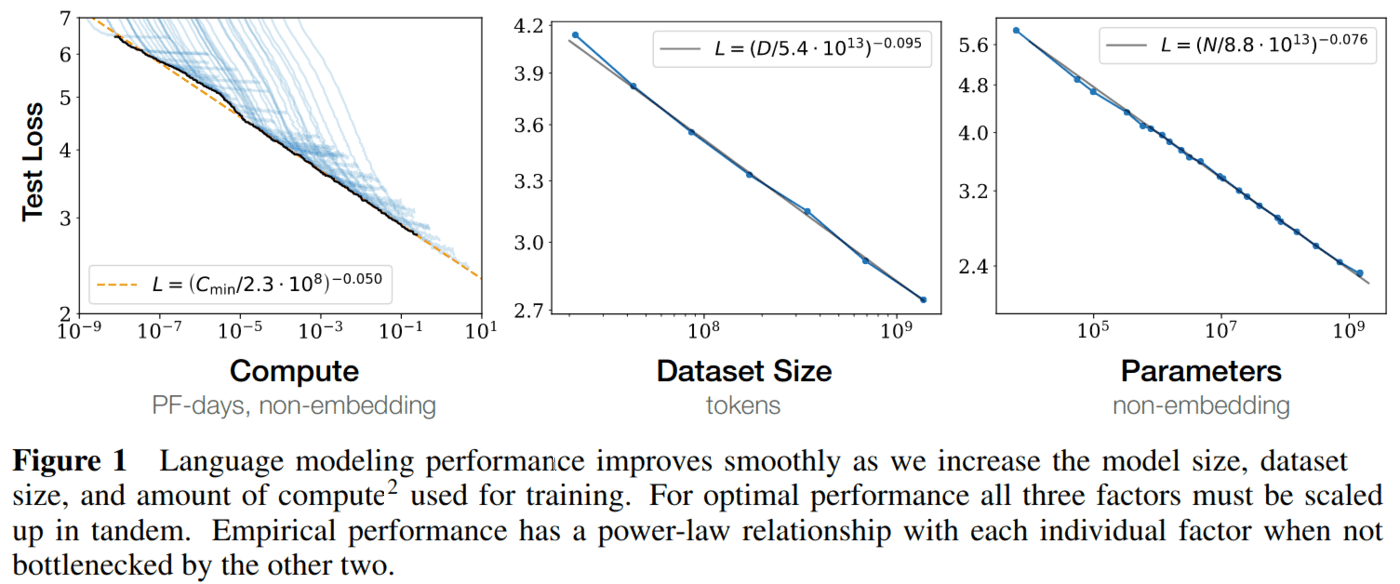
If this looks like hyperscaling plots that the AGI world is obsessed with atm, you are not wrong!
The difference is that rather than throwing tens or hundreds of $Billions at the problem, biomedicine is moving VERY SLOWLY toward sufficiently large datasets. Attention EA folks!

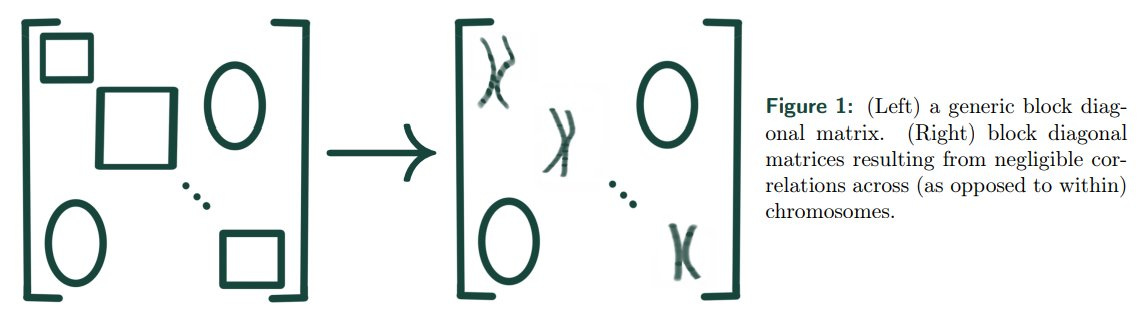
Hi Steve, nice paper! Would it be possible to identify and use sub-chromosome high LD blocks, or maybe overlapping smaller chunks done in parallel?